Scope: Treatment Effect Heterogeneity methodology - from traditional subgroup assessments to causal inference based on machine learning.
We are: Björn Bornkamp, Mathias Cardner, Chuyu Deng, Paulo Eusebi, Christine Fletcher, Jared Huling, Ilya Lipkovich, Karl Koechert, Nicole Kraemer, Henrik Loft, Brian Millen, Esha Mohammed, Heiko Goette, Huang Xin, Arne Ring, Bohdana Ratitch, Gerd Rosenkranz, Barbara Rosettani, Kostas Sechidis, Amy Spencer, David Svensson, Julien Tanniou, Marius Thomas, and Xu Yuejia,
The SIG lead is David Svensson (david.j.svensson@astrazeneca.com).
_______________________________________________________________________________________________________
Subgroup analysis is routinely conducted in drug development, in various settings; one key aspect is the regulatory requirement to demonstrate consistency of treatment effect across a pre-defined set of subgroups (e.g., ICHE5, E9, E17). This is performed as a risk-benefit assessment, aiming to identify the right patient population to treat - and, here, that set of subgroups is agreed with regulators prior to the trial conduct. Another key aspect is subgroup selection, where the aim is to estimate the effect in the most promising subpopulation (typically for planning another trial). The latter can either be done with respect to the same fixed set of pre-specified subgroups as mentioned earlier, or in a data driven fashion (e.g., biomarker subgroup detection).
There are well-known inherent statistical difficulties with all the above; with consistency, due to limited data in the subgroups, there is a high risk of false positives (random highs) as well as a low power to detect true differential effects (since trials are seldomly sized for it). In the subgroup selection setting, it is of key importance to provide an honest estimate discounted for the number of subgroups inspected, in order to not overstate the real effect. Even in a consistency assessment setting, there might well be a certain tendency to focus on the most deviating subgroup results, hence possibly introducing a bias although not formally a 'selection' problem.
The PSI Subgroup SIG submitted a White Paper in May 2018 on some of these aspects, containing an overview of the inherent problems, recommendations for the planning stage, a novel permutation based approach for assessing expected deviations under a null assumption, and some simulation based conclusions where various methods were compared. Due to the complexity not all available methods were initially studied (e.g., the Bayesian ones) and further work is being conducted. The aim is to provide an updated document later when these methods have been developed and evaluated.
Scope since 2018: the SIG is not only devoted to questions related to fixed (pre-specified) sets of subgroups, but also to the developments in personalized medicine, where lots of progress has been made in recent years on data-driven subgroup-detection/ML methodologies (causal inference, individual treatment effects and individual treatment regimes). The SIG aims to provide as much guidance and clarity as possible on inherent issues and approaches to the problems encountered in the these areas, and to promote cross industry research collaborations.
Some examples of topics discussed are: permutation-based approaches for assessing expected ranges and for generating NULL data/t1e control; Shrinkage methods of various kinds; Model averaging approaches; Bootstrap bias reduction, novel Graphical methods, general principles to consider in ML Subgroup Detection, ML in dose-response settings, ITR, causal inference.
BELOW: some material (including some presentations) which gives a flavour of the scope of the SIG.

Topics Discussed During Previous Meetings:
- 2018 APRIL: Update re progress of White Paper, remaining work for 2018. Ideas include further work on Bayesian shrinkage, Model averaging, Simulations under NULL when prognostic factors are present, SEAMOS development for non-linear models and Bootstrap Bias reduction.
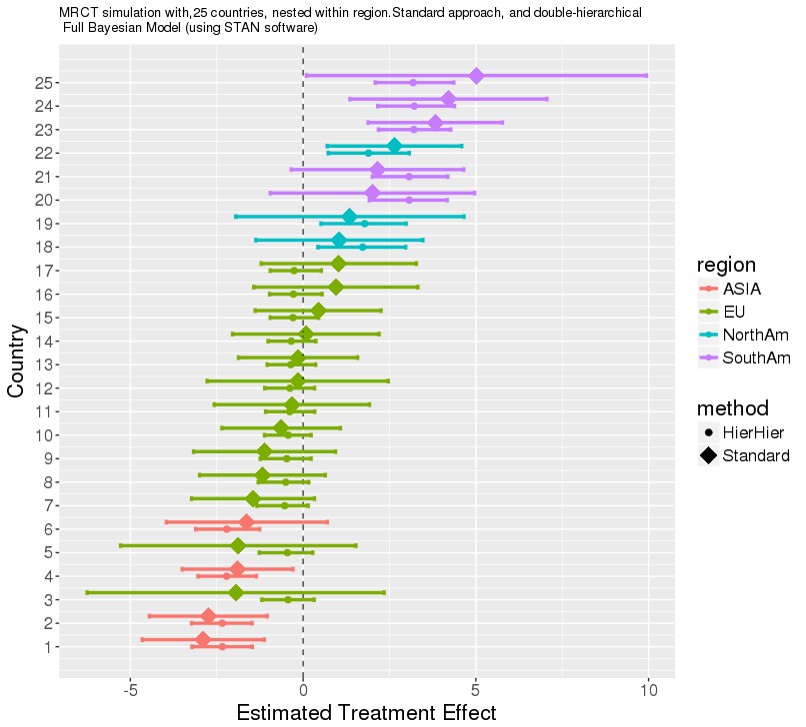
- 2018 JUNE: David PSI presentation on some aspects of Shrinkage, multi-level hierarchical models, and model averaging. Key aspect: many variations exist, and some unknowns regarding performance.
- 2018 JULY: Simulation of RCT discussed with underlying prognostic predictive continouos variables - dichotomized into subgroup factors - some preliminary illustrations of methods listed under APRIL.
- 2018 SEPT: Further simulations on the performance of BIC model averaging.
- 2018 NOV: Visualisation using novel R package SubrPlots (e.g., UpSet graph), Amy Spencer presenting further work on SEAMOS (modification to increase power). Updates on PSI 2019 Subgroup Section
- 2019 JAN: Discussing content for SIG subgroup session at PSI, presentation by David Svensson on SEAMOS (some simulation results in a non-linear case with prognostic factors).
- 2019 MARCH: Review of novel R CRAN package 'subtee', containing functionality for Bootstrap-based bias-reduction and BIC-Model-Averaging Shrinkage. Some simulation examples were discussed, and it was decided that more such (across broader assumptions) will be the topic for another SIG meeting. Issues regarding hierarchical Bayesian shrinkage when subgroups are overlapping was also discussed, with particular attention to a paper by Varadhan and Wang.
- EFSPI WORKSHOP 20th MARCH 2019 on Recent Developments on Subgroups and Biomarkers (Gothenburg) saw contributions from the SIG (presentations by G. Rosenkranz, and I.Lipkovich & D.Svensson) with the focus on HTE/Individual Treatment Effects, but also general aspects of consistency across subgroups was discussed, including some regulatory contribution.
- MAY 2019: Discussion of resampling principles for creating NULL data (idea: generating a large number of replicates with marginal distribution and correlations as in observed data except that treatment hetereogeneity is taken out); discussings also regarding possibities of a new points-to-consider document for exploratory analyses ('reality check' aspects to consider when assessing a given datamining/subgroup detection analysis).
- JUNE (PSI LONDON 2019): SIG members presented an overview and some recommendations regarding subgroup detection (Necdet Gunsoy, Ilya Lipkovich & David Svensson) at the SIG session (chaired by Aaron Dane).
- AUGUST 2019: Further discussions and planning of compiling a points to consider document that would help upfront when planning/conducting/reviewing an exploratory subgroup analysis (the latter to be interpreted in a broad sense, including data-driven approaches). Discussions regarding the upcoming PSI webex and PSI Subgroup session for Barcelona 2020.
- SEPTEMBER 2019: discussing best practices when performing ML SubId, initiating a long term SIG work in this direction. Common difficulties, common likely errors and how to avoid it.
- NOVEMBER 2019: further work on the above, planning a joint PSI talk in this direction.
- DECEMBER 2019: Further work on the joint presentation & possible SIG white paper.
- JANUARY 2020: on simulations for illustrating issues to avoid/overcome in SubId in the practice
- MARCH 2020: On formal Type 1 Error control in ML - novel ideas
- MAY 2020: Presentation and group discussions of a novel idea for SubId in Dose Response (paper by Marius Thomas, Bjoen Bornkamp, Heidi Seibold).
- JULY 2020: overview of recent ideas in the ITR area; optimizing Treatment for a given patient.
- OCTOBER 2020: ITR analyses in Multi-Treatment trials.
- NOVEMBER 2020: Knock-Offs Variable Importance (Kostas Sechidis)
- JANUARY 2021: Virtual Twins implementation aspects (shared surface, XGboost) and Causal Forest (David Svensson)
- MARCH 2021: Assessment Evaluation of different analytic strategies for estimating optimal treatment regimens for time-to-event outcomes in observational data (Ilya Lipkovich)
- 6th JUNE 2021: Presentation by Oliver Keene on Subgroup Dynamic Borrowing.
- 22th JUNE 2021: SIG SESSION at the PSI Conference (speakers: D. Svensson, D. Lawrance)
- 14th OCTOBER 2021: Discussing simulations regarding SEAMOS vs classical methods, beyond linear models (M. O'Kelly, D. Svensson)
- 17th NOVEMBER 2021: PSI SIG WEBINAR: Modern approaches to subgroup identification. (speakers: Svensson, Lipkovich, Bornkamp, Sechidis, Eusebi).
- 2nd DEC 2021: Discussing role and design of simulations for evaluation of methodologies.
- February 2022: group discussions of some recent papers (Modified covariate methods, value based for survival data)
- March: discussing BART; Bayesian Causal Forest (D. Svensson)
- May: Yuejia presented her Ph.D Thesis re optimal ITR in multi-arm settings
- June, 13: PSI Gothenburg SIG session; speakers David Svensson, Björn Bornkamp and Stefan Franzen.
- September, 26: SIG meeting covering PSI 2022, and a potential 2023 session, and ideas for collaborations.
- October 2022: Daniel Bratton - presentation of new ideas re Shrinkage without exchangeability assumptions
- November 2022: Nicole Kraemer - presentation of a resampling based approach for Consistency assessment
- December 16th 2022: Biomarker SIG Co-Leads visiting Subgroup SIG - connecting/sharing ideas
- JUNE 2023: PSI SIG session (speakers: Paulo Eusebi, Nicole Kraemer, Yuejia Xu; all co-work within the SIG)